General Overview
Mycobacterium avium subsp. paratuberculosis (MAP) is an obligate pathogenic bacterium of the genus Mycobacterium which causes chronic inflammation of the intestine in domestic and wild ruminants as well as other animals, including primates. M. avium subsp.paratuberculosis can live in animals for years without necessarily causing clinical disease. Infection is widespread in domestic livestock in Europe and North America but can occur elsewhere. [1]
Summary Data
O’Brien et al.(1976) exposed three groups of newly weaned 4-month-old red deer orally with M. avium subsp. paratuberculosis Bovine strain and necropsy was conducted 44 weeks post inoculation to determine the infection rate.[2]
Brotherston et al. (1976) inoculated South Country Cheviots at the age of three weeks orally with M. avium subsp. paratuberculosis IOI strain which was originally recovered from a clinical case of the disease in a sheep and VB/4 strain from an affected cow.[3] The necropsy was done 1-9 months post inoculation.
Recommended Model
Experiment number 262 model is the recommended model. The ID50 of the model has lower value than the other model.
References
- (2005). Mycobacterium avium Subspecies paratuberculosis Cultured from Locally and Commercially Pasteurized Cow's Milk in the Czech Republic. Applied and Environmental Microbiology. 71, 1210-1214.
- (2006). Immunological and molecular characterization of susceptibility in relationship to bacterial strain differences in Mycobacterium avium subsp. paratuberculosis infection in the red deer (Cervus elaphus). Infection and Immunity. 74, 3530.
- (1961). Quantitative studies of Mycobacterium johnei in the tissues of sheep. Journal of Comparative Pathology. 71, 286.
| ID | # of Doses | Agent Strain | Dose Units | Host type | Μodel | Optimized parameters | Response type | Reference |
|---|---|---|---|---|---|---|---|---|
| 262 | 3 | sub sp. Paratuberculosis Bovine | CFU | deer | exponential |
k = 6.93E-04 LD50/ID50 = 1000 |
infection | "Quantitative studies of Mycobacterium johnei in tissues of sheep. III. Intestinal histopathology." Journal of comparative pathology. 72 (1962): 80. |
| 263 | 3 | sub sp. Paratuberculosis IOI strain | CFU | cheviots | beta-Poisson |
a = 5.79E-02 LD50/ID50 = 4.8E+02 N50 = 4.8E+02 |
infection | "Pathogenicity of Acanthamoeba and Corynebacterium in the Rat Cornea." Archives of Ophthalmology. 108 (1990): 1. |
| myco_intravenous | 5 | M. avium serovar 1, human non- AIDS isolates: pooled strains N-289, N-356, N-357, N-364, N-445, N458, N-461; Serovar 9 strains N-254, N-302 | CFU | mice | exponential |
k = 1.10E-6 LD50/ID50 = 6.30E+5 |
Lung lesions | |
| myco_intravenous_2 | 5 | MAC 101 (human-derived) | CFU | mice | exponential |
k = 3.12E-9 LD50/ID50 = 2.22E+8 |
death | |
| myco_intravenous_3 | 15 | M. avium serotype II strain SSC 1323 (porcine-derived | CFU | pig | exponential |
k = 9.26E-9 LD50/ID50 = 7.49E+7 |
Lesions or culture in lymph nodes | |
| myco_intravenous_3 | 15 | M. avium serotype II strain SSC 1323 (porcine-derived) | CFU | pig | exponential |
k = 7.88E-8 LD50/ID50 = 8.79E+6 |
Lesions or culture in lymph nodes | |
| myco_intravenous_4 | 15 | CFU | pig | exponential |
k = 1.65E-8 LD50/ID50 = 4.19E+7 |
Lesions or culture in lymph nodes | ||
| myco_intratracheal | 3 | MAC serotype 8 (Human-derived, AIDS Patient) | CFU | hamsters | beta-Poisson |
a = 0.201 k = 4.87E-9 LD50/ID50 = 1.42E+8 N50 = 1.42E+8 |
Disseminated infection | |
| myco_intratracheal_2 | 3 | MAC serotype 8 (Human-derived, AIDS Patient) | CFU | hamsters | beta-Poisson |
a = 0.048 LD50/ID50 = 7.93E+8 N50 = 7.93E+8 |
Liver infection |
k = 6.93E-04
LD50/ID50 = 1000
|
|
||||||||||||||||||||||
|
||||||||||||||||||||||||||||||||||||||
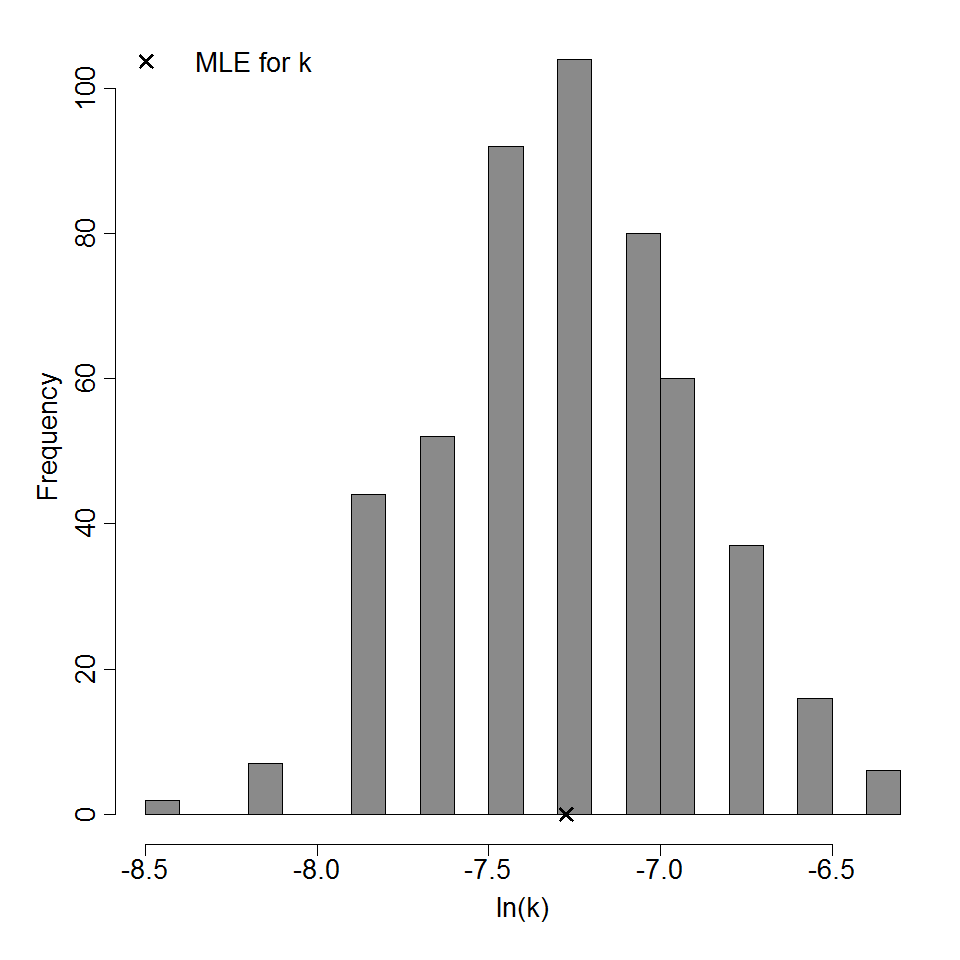
Parameter histogram for Exponential model (uncertainty of the parameter)

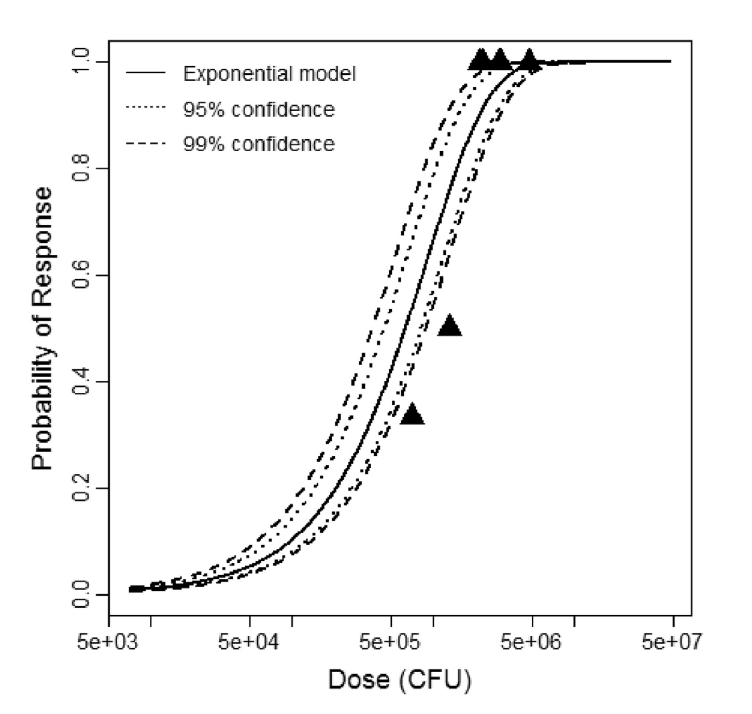
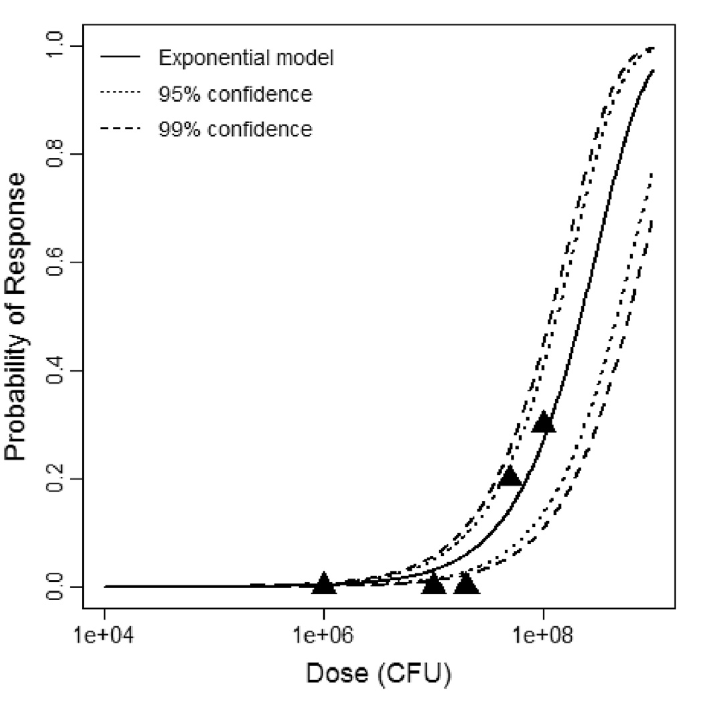
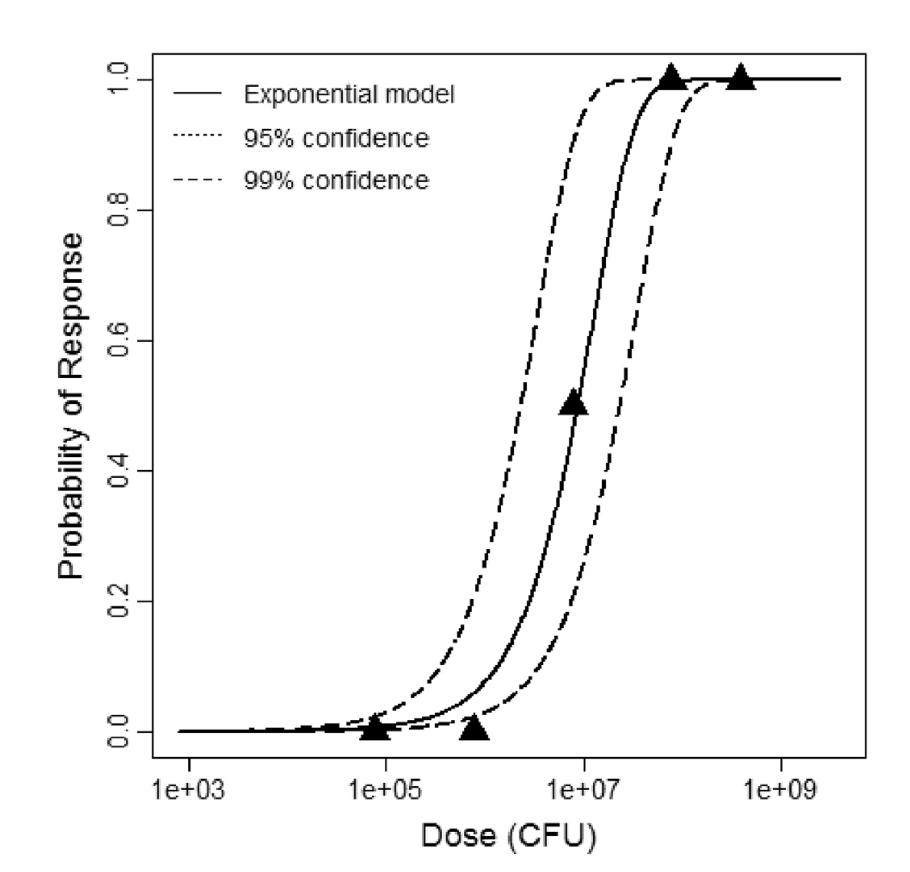
Exponential model plot, with confidence bounds around optimized model
References
LD50/ID50 = 4.8E+02
N50 = 4.8E+02
|
|
||||||||||||||||||||||
|
||||||||||||||||||||||||||||||

Parameter scatter plot for beta Poisson model ellipses signify the 0.9, 0.95 and 0.99 confidence of the parameters

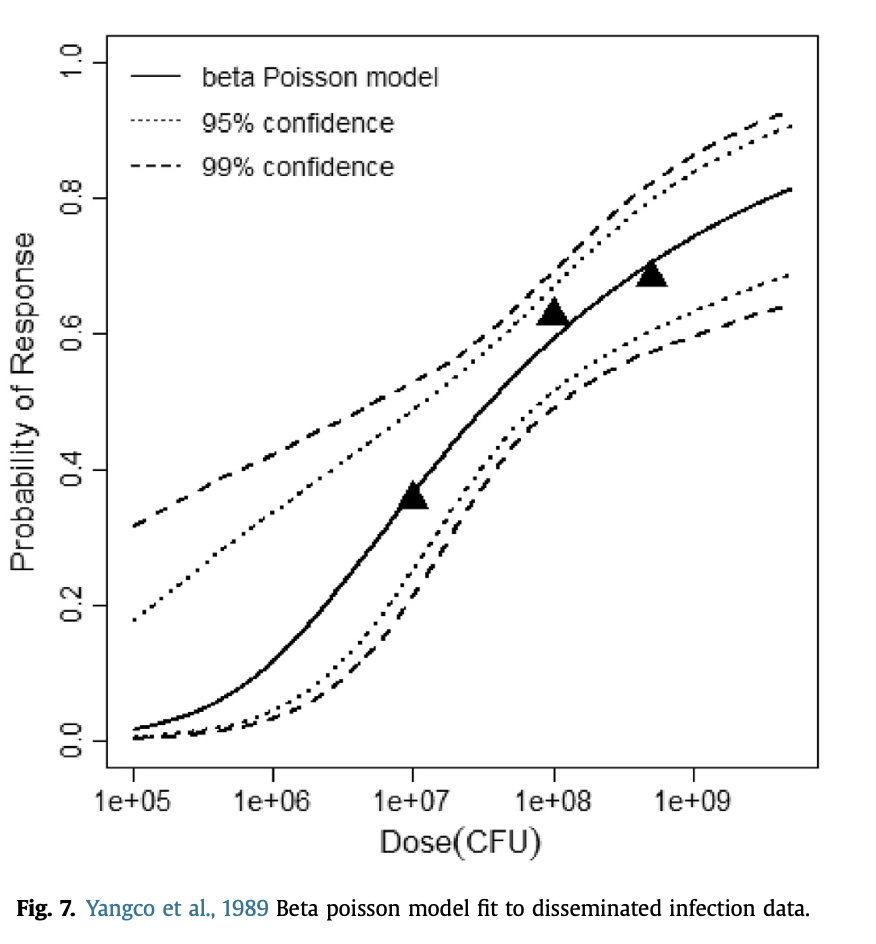
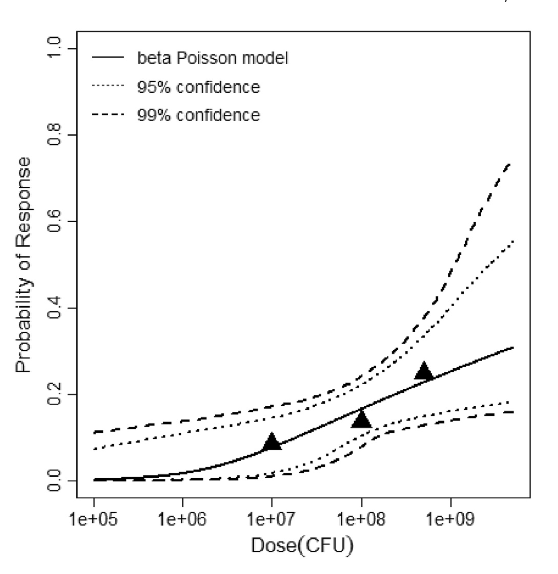
beta Poisson model plot, with confidence bounds around optimized model
References
k = 1.10E-6
LD50/ID50 = 6.30E+5
k = 3.12E-9
LD50/ID50 = 2.22E+8
k = 9.26E-9
LD50/ID50 = 7.49E+7
k = 7.88E-8
LD50/ID50 = 8.79E+6
k = 1.65E-8
LD50/ID50 = 4.19E+7
k = 4.87E-9
LD50/ID50 = 1.42E+8
N50 = 1.42E+8
LD50/ID50 = 7.93E+8
N50 = 7.93E+8
 QMRA
QMRA