General Overview
Salmonella is a genus of rod-shaped, gram-negative, non-spore forming, predominantly motile enterobacteria [1]. Many species of Salmonella have been isolated from eggs and egg products [2]. Salmonella meleagridis is one of the most common serotype of Salmonella [3]. Twenty human isolates of S. meleagridis had been identified in Canada so far during 1997.
http://www.cdc.gov/salmonella/
Summary Data
McCullough, and Eisele (1951) inoculated human volunteers orally with the S. Meleagridis strain I,II and III.[4]
Recommended Model
Eexperiment number 238 is the recommended one as the strain I was more virulent than strain III.
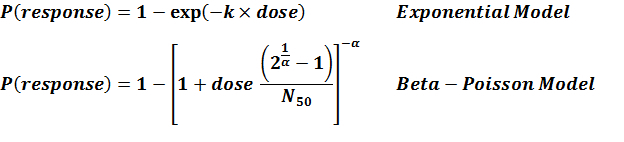
References
- (1988). A review of human salmonellosis: III. Magnitude of Salmonella infection in the United States. Reviews of infectious diseases. 10, 111-124.
- (1951). Experimental human salmonellosis: I. Pathogenicity of strains of Salmonella meleagridis and Salmonella anatum obtained from spray-dried whole egg. Oxford Journal of Infectious Diseases. 88(3),
- Citekey 1275 not found
- (1951). Experimental Human Salmonellosis: I. Pathogenicity of Strains of Salmonella Meleagridis and Salmonella Anatum Obtained from Spray-Dried Whole Egg. Oxford Journal of Infectious Diseases. 88(3),
| ID | # of Doses | Agent Strain | Dose Units | Host type | Μodel | Optimized parameters | Response type | Reference |
|---|---|---|---|---|---|---|---|---|
| 238 | 11 | strain I | CFU | human | beta-Poisson |
a = 3.89E-01 LD50/ID50 = 1.68E+04 N50 = 1.68E+04 |
infection | "Experimental Human Salmonellosis: I. Pathogenicity of Strains of Salmonella Meleagridis and Salmonella Anatum Obtained from Spray-Dried Whole Egg." Oxford Journal of Infectious Diseases. 88.3 (1951). |
| 240 | 4 | strain III | CFU | human | beta-Poisson |
a = 8.85E-01 LD50/ID50 = 5.24E+05 N50 = 5.24E+05 |
infection | "Changes in Iron and Transferrin Levels and Body Temperature in Experimental Airborne Legionellosis." The Journal of Infectious Diseases. 147 (1983): 2. |
LD50/ID50 = 1.68E+04
N50 = 1.68E+04
|
|
||||||||||||||||||||||
|
||||||||||||||||||||||||||||||
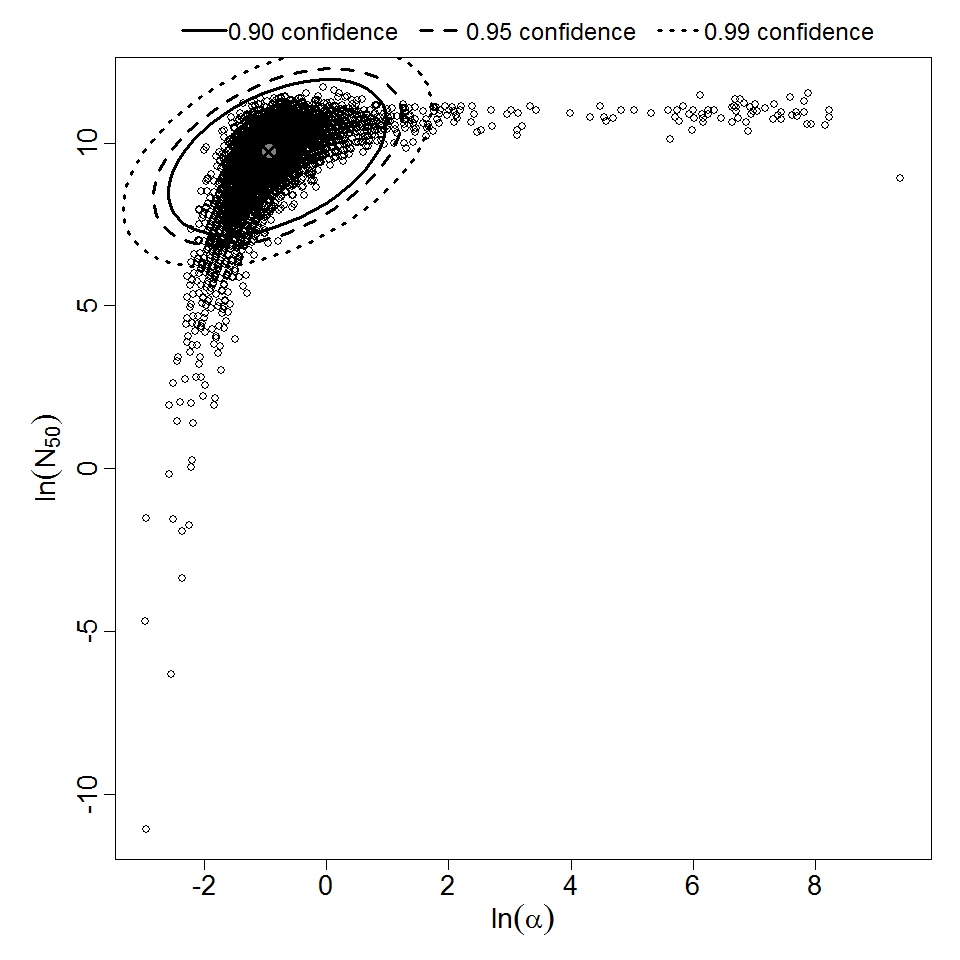
Parameter scatter plot for beta Poisson model ellipses signify the 0.9, 0.95 and 0.99 confidence of the parameters.
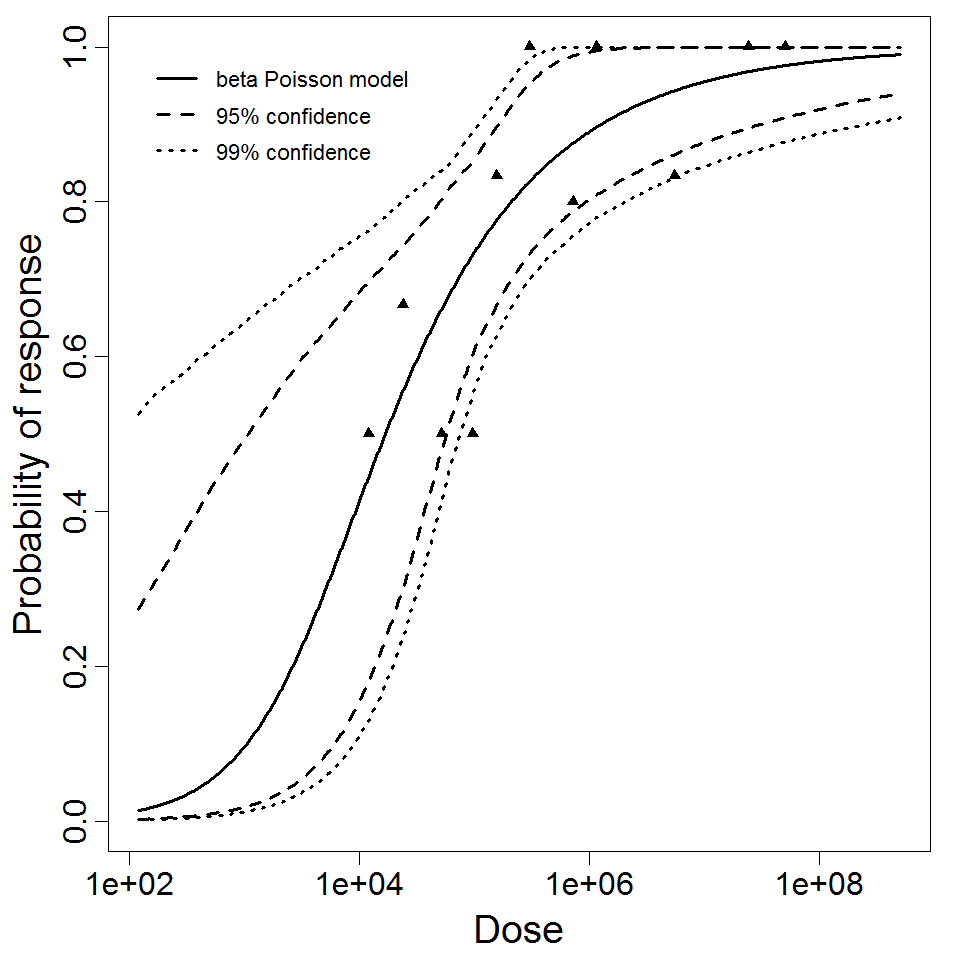
beta Poisson model plot, with confidence bounds around optimized model
References
LD50/ID50 = 5.24E+05
N50 = 5.24E+05
|
|
||||||||||||||||||||||
|
||||||||||||||||||||||||||||||
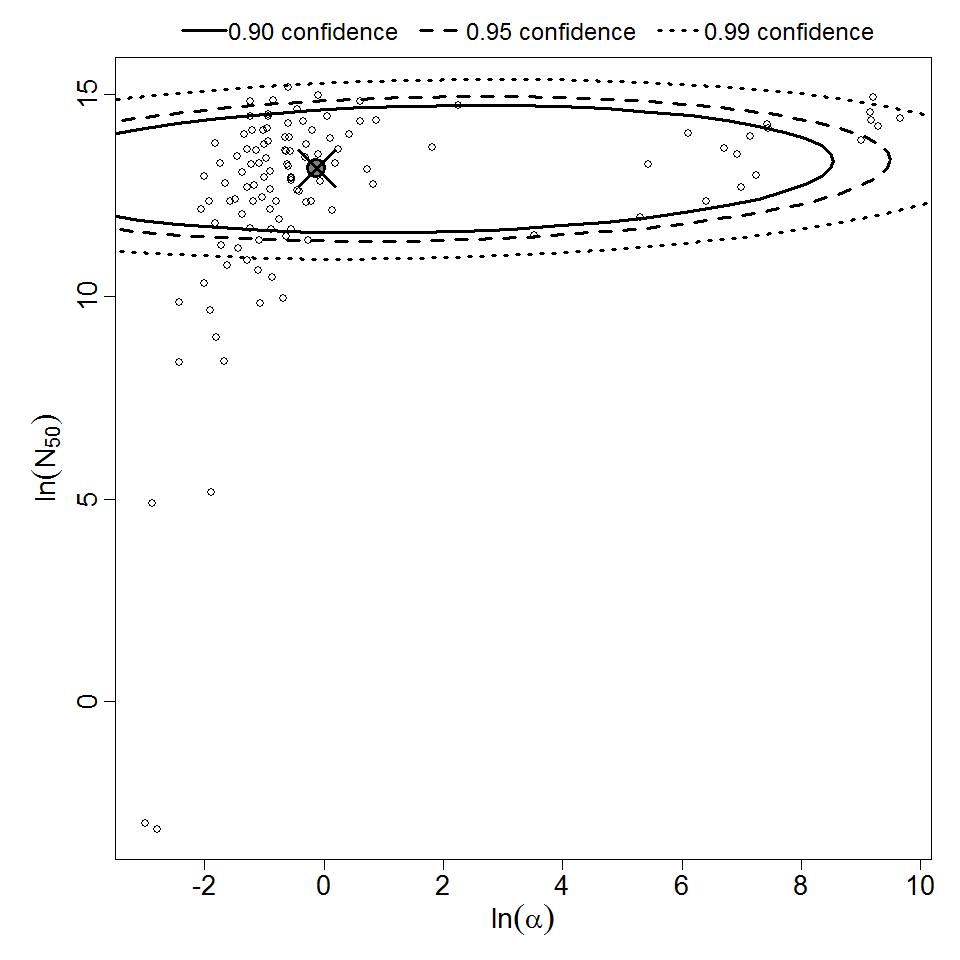
Parameter scatter plot for beta Poisson model ellipses signify the 0.9, 0.95 and 0.99 confidence of the parameters.
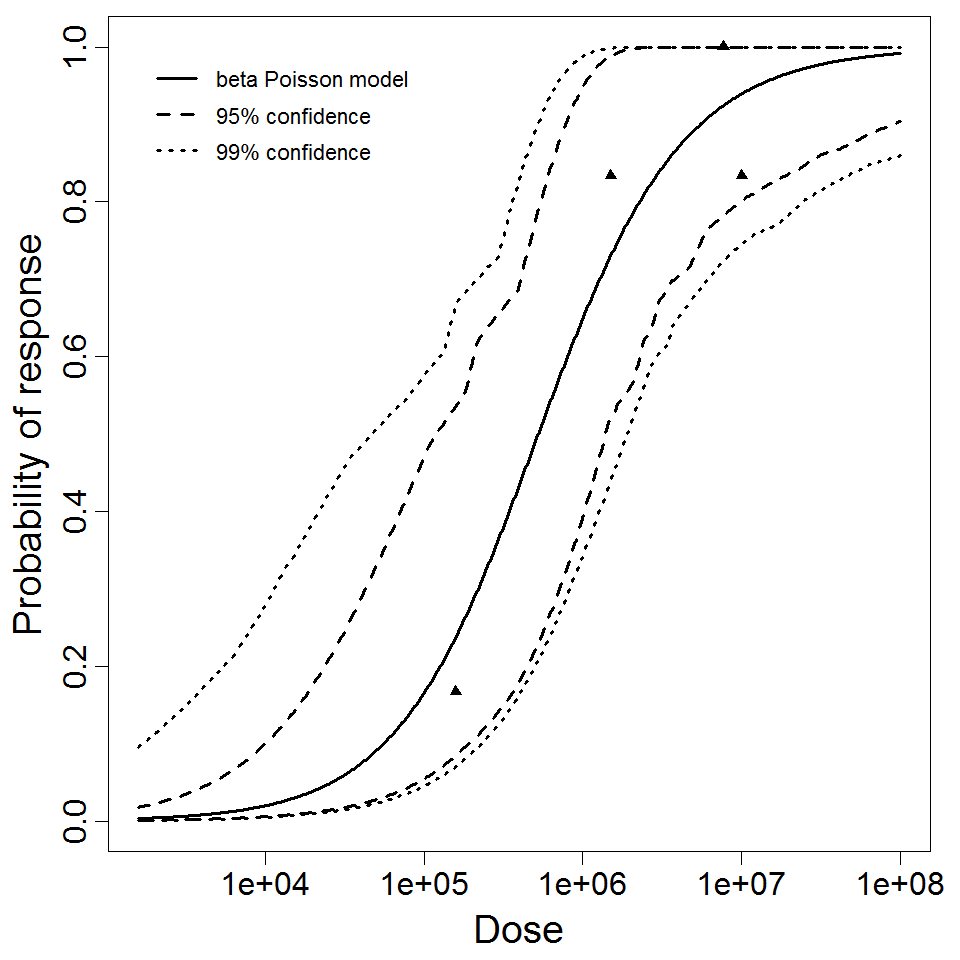
beta Poisson model plot, with confidence bounds around optimized model
 QMRA
QMRA