Author: Jade Mitchell and Sushil B. Tamrakar
General overview
Inactivation of vegetative microorganisms, there are several types of survival curves that described the inactivation rates and patterns(Xiong, Xie et al. 1999)[1]. The most commonly used mathematical linear model is first order exponential model. However, in many cases linear model alone cannot describe the prevailing pattern. There are several nonlinear models described by various investigators to fit the data (Coroller, Leguerinel et al. 2006)[2]. Seven different models described in various studies have been shown in Table 1.(Peleg and Cole 1998[3]; Juneja, Eblen et al. 2001[4]; Valdramidis, Bernaerts et al. 2005[5]; Juneja, Huang et al. 2006[6]).
The data from each treatment was fitted to a best-fit curve using an unpublished mathematical model fitting tool in Microsoft® Excel (Microsoft® Inc., Redmond, Washington) by Patrick Gurian at Drexel University and modified by Sushil Tamrakar (Michigan State University). The tool can be used to model bacterial survival in culture-dependent or culture-independent methods independent of the organism and the environmental conditions. The smallest absolute value of the Bayesian information criterion (BIC) was the criteria to choose the best fit model.
Table 1 Persistent models and equations
|
S.N. |
Model |
Equation |
Curve Properties |
Reference |
|
1 |
First order exponential decay model |
Ln (Nt/N0) = -kt |
Linear, negative slope |
Crane and Moore, 1986 |
|
2 |
Biphasic exponential decay model |
for 0≤t<x: Ln (Nt/N0) = -k1*t for t≥x: Ln (Nt/N0) = -k1*t+k2*(t-x) |
Linear, negative slope, slope changes at t=x |
Carret et al. 1991 |
|
3 |
General logistic model |
Ln (Nt/N0) = ln(2/(1+e(-kt))) |
Nonlinear, concave |
Gonzalez, 1995 |
|
4 |
Exponential damped model |
Ln (Nt/N0) = -kt*e(-st) |
Nonlinear, concave |
Cavalli-Sforza et al. 1983 |
|
5 |
Gompertz model |
Ln (Nt/N0) = ln[C*e [-e^(-b*(ln(t)-a))]} |
Nonlinear, concave |
Gompertz, 1825 Gil et al. 2011[7] |
|
6 |
Two-stage model Juneja and Marks (1) |
Ln (Nt/N0) = -Ln(1-(1-e(-kt)^m))) |
Nonlinear, concave |
Juneja et al. 2006 |
|
7 |
Log-logistic model Juneja and Marks (2) |
Ln (Nt/N0) = -Ln(1+e (a+b*ln (t))) |
Convex or concave |
Juneja et al. 2003 |
|
8 |
Gompertz-3 (gz3) |
Ln (Nt/N0) =(k1*exp(-exp((-k2*exp(1)*(k3-t)/k1)+1) ) ) |
k1, k2, k3 > 0 |
Gil et al. 2011[7] |
|
9 |
Gompertz-Makeham (gzm) |
Ln (Nt/N0)=(-k3*t -k1/k2*(exp^((k2*t)).-1)) |
k1, k2, k3 > 0 |
Jodra, 2009[8] |
|
10 |
Weibull (wb) |
Ln (Nt/N0)=-{(t/k1)}^k2 |
k1= treatment time for first decimal reduction |
|
|
11 |
Gamma (gam) |
Ln (Nt/N0)=(t^(k2-1) ) exp^((-t/k2)). |
Nonlinear concave |
[1],[2] |
|
12 |
Sigmoid-B (sB) |
Ln (Nt/N0)=-(k1*t^k3)/(k2+t^k3 ) |
Nonlinear concave or convex |
Peleg 2006[10] |
|
13 |
Logistic-Fermi Combination |
N(t)/N_0 =1/(1+exp(k1*(t-k2))) |
Nonlinear, concave |
Peleg 2006[10] |
|
14 |
Log-normal |
N(t)/N_0)=[1-Φ{(ln(t)-µ)/б}] |
Concave, nonlinear |
Aragao 2007. |
|
15 |
Biphasic(3 parameters) |
if (min(t)<k3) |
Nonlinear, concave |
Kamau et al. (1990) |
|
16 |
Double exponential |
N(t)/N_0 =a exp(-k1t) + (1-a) exp(-k2t) |
Nonlinear, concave |
Peleg 2006[10] |
|
17 |
Sigmoid type A |
log(N(t)/N_0 )=(a_1 t)/([1+a_2 t][a_3-t]) |
Nonlinear, S curve |
Peleg 2006[10] |
The data from fomite recovery experiments in Dr. Charles Gerba’s lab were fit to the different persistent models. The best fit models are shown in Table 2 and Figure 1 .
Table 2 Best-fit models
|
Organism |
Fomite |
Time(hrs) |
Best fit model |
|
B. anthracis |
Polyester |
0,24,672,2190 |
Biphasic exponential |
|
B. anthracis |
Steel |
0,24,672,2190 |
Biphasic exponential |
|
B. anthracis |
Laminar |
0,24,672,2190 |
Juneja & Mark(2) |
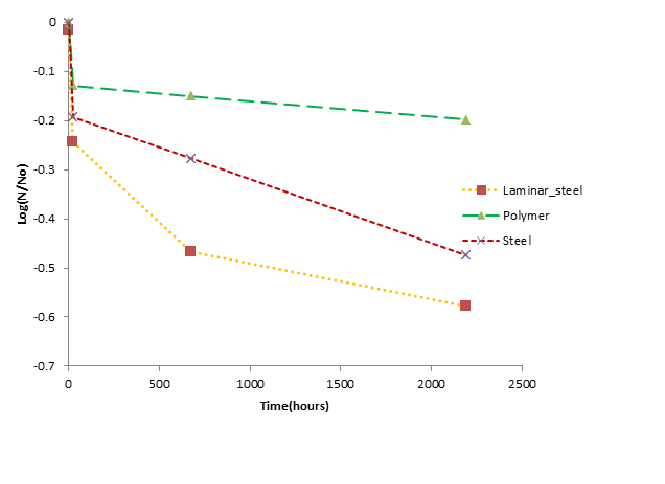
Figure 1 Best fit models
Persistence excel tool
Any persistent data could be analyzed by using attached excel spreadsheet. Read the instruction manual first and then use the excel tool accordingly.
References
- (1999). A mathematical model for bacterial inactivation. International journal of food microbiology. 46, 45–55.
- (2006). General model, based on two mixed Weibull distributions of bacterial resistance, for describing various shapes of inactivation curves. Applied and Environmental Microbiology. 72, 6493–6502.
- (1998). Reinterpretation of microbial survival curves. Critical Reviews in Food Science. 38, 353–380.
- (2001). Modeling non-linear survival curves to calculate thermal inactivation of Salmonella in poultry of different fat levels. International journal of food microbiology. 70, 37–51.
- (2005). An alternative approach to non-log-linear thermal microbial inactivation: modelling the number of log cycles reduction with respect to temperature. Food Technology and Biotechnology. 43, 321–327.
- (2006). Predictive model for growth of Clostridium perfringens in cooked cured pork. International journal of food microbiology. 110, 85–92.
- (2011). On the use of the Gompertz model to predict microbial thermal inactivation under isothermal and non-isothermal conditions. Food Engineering Reviews. 3, 17–25.
- (2009). A closed-form expression for the quantile function of the Gompertz–Makeham distribution. Mathematics and Computers in Simulation. 79, 3069–3075.
- (2002). On calculating sterility in thermal preservation methods: application of the Weibull frequency distribution model. International journal of food microbiology. 72, 107–113.
- (2006). Advanced quantitative microbiology for foods and biosystems: models for predicting growth and inactivation.
 QMRA
QMRA